 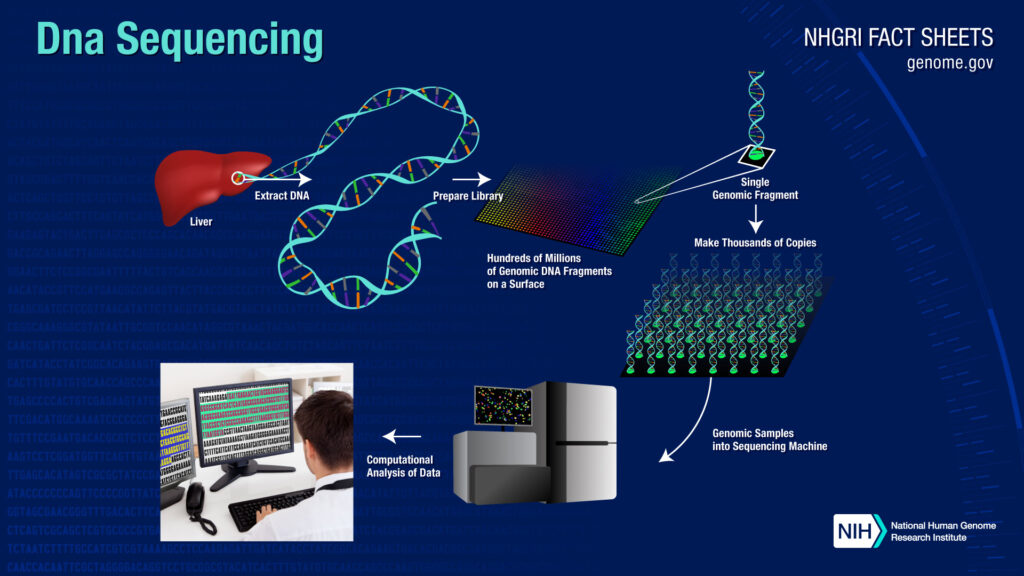
Next-generation sequencing technology brings the opportunity for robust and efficient characterization of the whole genome of any pathogenic organism at any point in time (Deurenberg et al. The production of molecular signature tools can only be possible after quick and complete molecular characterization of any pathogen. On the other hand, generation of molecular signatures for the pathogens will probably be the answer for every problem. The repetition of sequence experiment with each sample and performing molecular analyses every time also needs some expertise from the laboratory personnel. Any novel pathogenic species is not only very much difficult to identify through this technique but characterization, as well as probable prevention strategy determination, also become a big problem. This technique can identify the pathogen (both culturable and non-culturable) correctly but it needs some specific background information to perform the experiment. The sequence data is then matched with available rDNA sequence data using a different molecular analysis tool for the accurate detection of the pathogen (John et al. The prokaryotic microorganisms are characterized through 16 s rDNA sequencing and characterization of eukaryotic microorganisms are done through 18 s rDNA sequencing technology (Deurenberg et al. During the last two decades, molecular characterization of the pathogen is customarily done through ribosomal DNA (rDNA) sequencing methods. For this problem, molecular diagnostics become increasingly popular day by day (Taylor et al. The pathogenic variability and co-evolutionary modifications of the microorganisms evade many pathogens from successful identification in many cases (Deurenberg et al. Although kit-based methods are quick, it demands more accuracy in many cases. Many molecular and biochemical test kits are also available in the market for the quick identification of the pathogen. These culture methods are time-consuming as different disease-causing microorganisms demand varied incubation times for their required growth. The identification of the pathogen is solely dependent on culture methods in diagnostic laboratories. Initial diagnosis of any infectious disease largely depends upon successful, proper as well as quick detection of the pathogen (Stafford et al. The infectious disease can be caused by a virus, bacteria, fungi, algae as well as many other eukaryotic microscopic organisms too (Daszak et al. 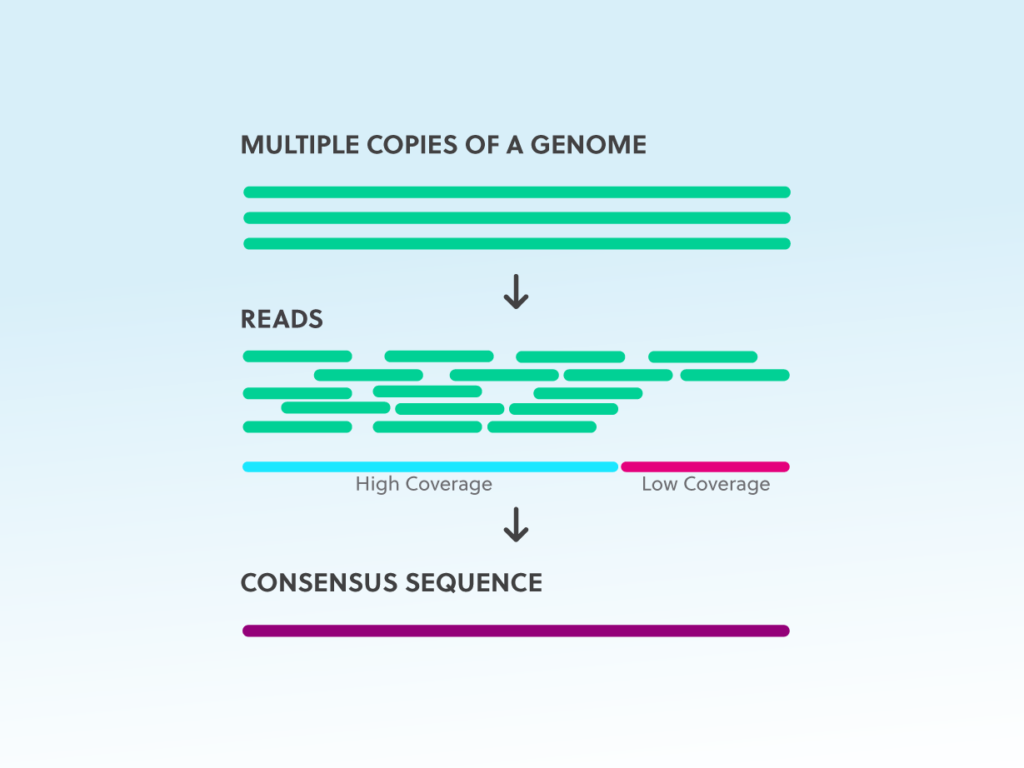
Medical microbiology is an important field of research since the past, which deals with the identification, characterization, and prevention of any disease-causing microorganisms. Therefore, the present review is focused on a brief introduction to NGS technology, its application in medical microbiology, and possible future aspects for the development of medical sciences. Metagenomic next generation sequencing is another specialized type of NGS that is profoundly utilized in medical biotechnology and diagnostics now a days. Recently, with the advent of next-generation sequencing (NGS) technology identification, and characterization of almost all the pathogenic bacteria become possible without any information a priori. Molecular diagnostics became a popular alternative to classical techniques for the past couple of decades but it required some prior information to detect the pathogen successfully. Classical cultural, as well as biochemical test-based identification, has its own limitations to their time-consuming and ineffectiveness for closely related pathovars. This is a real challenge to the scientific as well as a medical community to find out a proper and robust method of pathogen detection. Rapid isolation, characterization, and identification are prerequisites of any successful medical intervention to infectious disease treatment.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
